What is Spark and Spark SQL?
Spark is an open source, scalable, massively parallel, an in-memory execution environment for running analytics applications. Think of it as an in-memory layer that sits above multiple data stores, where data can be loaded into memory and analyzed in parallel across a cluster. Spark SQL is a Spark module for structured data processing. It provides a programming abstraction called DataFrames and can also act as a distributed SQL query engine.
Note: dataset mentioned in the following discussion is not dataset API in spark. It’s a set of data.
There are N numbers of things responsible for bad or below average performance. I am going to touch base only things that I come across and how I addressed those issues.
When optimizing the performance of Spark-SQL or any other distributed computing framework it’s very very important to know your data (KYD) first. Because spark job may run exceptionally well with one set of data and really bad for other. We are going to address painful joins in the following scenarios
- Joins between datasets with different cardinalities
- Skewed data joins
How joins works?
Following diagram works how normally joins works in spark-SQL. This join is also known as shuffle join. Because when we join two datasets, let’s say A & B, data for one key is brought on one executor. (Small boxes inside rectangle show executors on different machines).
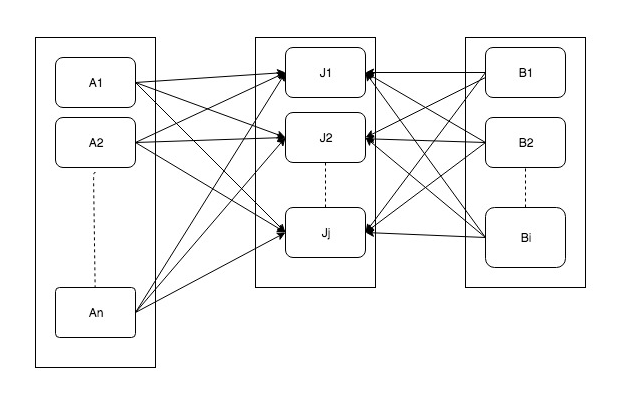
In some of the scenarios above operation can be problematic. In worst it may result in complete data movement of both datasets. This may result in network congestions and also increase I/O. Also if there is a lot of data against one key (skewed data) it can result in job failure or at least terribly slow down job execution. In the following topics, we will see how to solve these problems.
- a) Joins between datasets with different cardinalities:
Joins can be of two types. Map-side join and Reduce-side join. Both joins perform good or bad depending on the datasets.
1) Consider, there are two datasets A and B. When we join A & B in spark automatically reduce-side join happens. That means spark shuffle the data for similar keys on the same executor. This join does well when the distribution of the data is uniform across the join-keys (keys/fields on which joins are happening)
e.g. If I join df_A.join(df_B, [key1, key2]) distribution of records for both key1 and key2 in both sets play an important role. If records are distributed uniformly across all join keys in both datasets taking part in this join.
To improve the performance of reduce join we should join on the minimal datasets. Minimal can be obtained by filtering out unwanted records before joins instead of doing this after the join. This way we can avoid unnecessary shuffling of data, as records for only those keys will be shuffled which are actually needed in results.
2) Now consider A is big datasets and B is a really small dataset. Now, what qualifies for the small dataset? As per spark documentation, 2GB is max-threshold for the auto-broadcast parameter. I am kind of agree with this threshold because this data will be copied to each executor. Datasets which few MB to 2-3 GB in size must be broadcasted.
This number depends on how big are the executors in the cluster. Also, roughly size of dataset * number of executors should not bigger than the bigger dataset. Otherwise using simple reduce side join will be efficient. Following diagram shows how broadcast joins works. You can see on each executor where partition for A exists, entire B dataset is made available.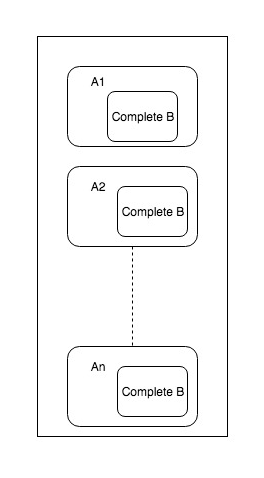
Enough theory!! How we can broadcast smaller dataset? It’s really simple. We just need to provide the broadcast hint to spark processing engine and that’s it. Following code snippet showcase how it can be done.
With both above approach check the shuffle size in spark-UI. Even if shuffle size is less, that does not mean amount of data-transferred is less. Might be in case of broadcast join we are moving the larger amount of data. Try out both to check performance and adopt which suits best for your need.
- b) Skewed data joins:
Consider we are joining dataset A and B. Skewed data means data with the un-uniform distribution of records across keys. More precisely huge %tage of data have very few numbers of keys. Now due spark processing the way joins are performed a large amount of data is collected on single or very few executors. This has following effects
- Tasks with large data run for a very very long time. Ultimately jobs take longer to finish. Other executors sit idle during this time. So ineffective usage of resources.
- If records for the key are too large, executors run out of memory and job fails.
Problem is explained/shown in the following diagram
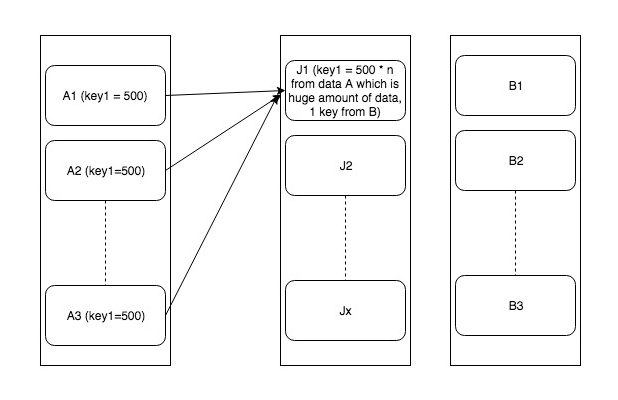
So one obvious solution of that is to remove these keys with skewed data. But that may not be possible every time. In such cases where it’s mandatory to process everything, we have two solutions
- Broadcast smaller dataset, if one of data is smaller
- If broadcasting is not possible, we have to split the data against skewed keys to more number of executors by adding the limited-range random number in join key in the dataset which has skewed records. Let’s call it A. But doing this will produce wrong results as records for skewed keys from other datasets (let’s call it B) may not be available at executors where we have distributed records from A.
To handle this we will multiply data in set B by cross join with limited range number dataset (lets 1-30 for distributing it to 30 records). Also, the random number within 1-30 range to each record in set A. Now join A and B on original join key “key1” and “random_val”. This will avoid executors from being flooded with a large number of keys. But be careful if we increase range 1-30 too much it will grow the data B exponentially. This is explained in the following diagram
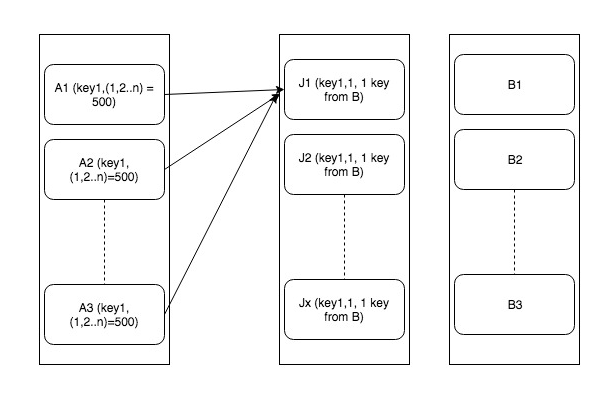
Here is how it can be done –
Conclusion:
- Broadcast joins are really helpful in the case of one small and one big dataset joins. It can drastically improve performance as well as network utilization.
- Datasets with skewed data can be handled efficiently by distributing data to different executors by adding the key to it. It can improve the performance and stability of the job run.









